Jane Flint - Principles of Virology
Здесь есть возможность читать онлайн «Jane Flint - Principles of Virology» — ознакомительный отрывок электронной книги совершенно бесплатно, а после прочтения отрывка купить полную версию. В некоторых случаях можно слушать аудио, скачать через торрент в формате fb2 и присутствует краткое содержание. Жанр: unrecognised, на английском языке. Описание произведения, (предисловие) а так же отзывы посетителей доступны на портале библиотеки ЛибКат.
- Название:Principles of Virology
- Автор:
- Жанр:
- Год:неизвестен
- ISBN:нет данных
- Рейтинг книги:3 / 5. Голосов: 1
-
Избранное:Добавить в избранное
- Отзывы:
-
Ваша оценка:
- 60
- 1
- 2
- 3
- 4
- 5
Principles of Virology: краткое содержание, описание и аннотация
Предлагаем к чтению аннотацию, описание, краткое содержание или предисловие (зависит от того, что написал сам автор книги «Principles of Virology»). Если вы не нашли необходимую информацию о книге — напишите в комментариях, мы постараемся отыскать её.
Volume I: Molecular Biology
Volume II: Pathogenesis and Control
Principles of Virology, Fifth Edition
Principles of Virology — читать онлайн ознакомительный отрывок
Ниже представлен текст книги, разбитый по страницам. Система сохранения места последней прочитанной страницы, позволяет с удобством читать онлайн бесплатно книгу «Principles of Virology», без необходимости каждый раз заново искать на чём Вы остановились. Поставьте закладку, и сможете в любой момент перейти на страницу, на которой закончили чтение.
Интервал:
Закладка:
One-step growth curve analysis can provide quantitative information about different virus-host systems. It is frequently employed to study mutant viruses to determine what parts of the infectious cycle are affected by a particular genetic lesion. It is also valuable for studying the multiplication of a new virus or viral reproduction in a new virus-host cell combination.
When cells are infected at a low multiplicity of infection, several cycles of viral reproduction may occur ( Fig. 2.18B). Growth curves established under these conditions can also provide useful information. When infection is carried out at a high multiplicity, a mutation may fail to have an obvious effect on viral reproduction. The defect may only become evident following a low-multiplicity infection. Because the effect of a mutation in each cycle is multiplied, a small effect can be amplified after several cycles. Defects in the ability of viruses to spread from cell to cell may also be revealed when multiple cycles of reproduction occur.
Global Analysis
The study of replication cycles of many viruses with one-step growth analysis has allowed a reductionist approach to understanding and defining the steps of virus attachment, entry, replication, and assembly. In contrast, new experimental and computational tools permit global analysis of viral, cellular, and host responses to infection. Global analyses apply a dizzying array of different high-throughput technologies to measure system-wide changes in DNA, RNA, proteins, and metabolites during virus infection of cells, tissues, or entire organisms. Data obtained from high-throughput measurements are integrated and analyzed using mathematical algorithms to generate models that are predictive of the system. For example, virus infections of different animals are characterized by the induction of distinct sets of cytokine genes, a property that can be correlated with different pathogenic outcomes. When a model has been developed, it can be further refined by the use of viral mutants or targeted inhibition of host genes or pathways. Global analysis is therefore a holistic, host-directed approach that complements traditional methods for studying viruses.
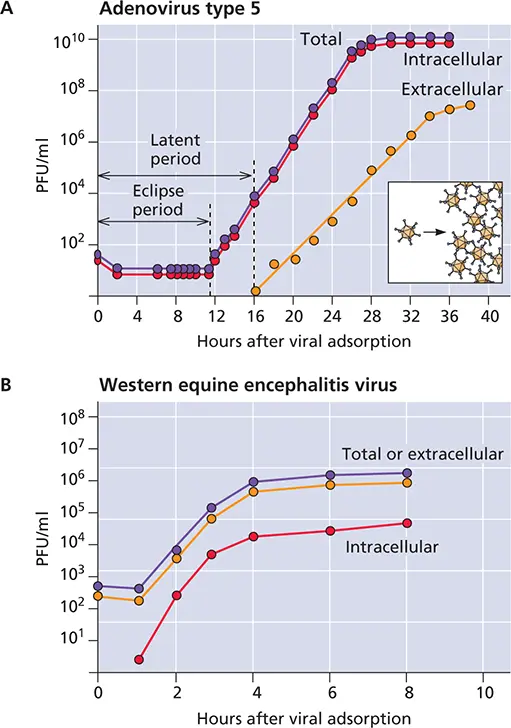
Figure 2.19 One-step growth curves of animal viruses. (A)Growth of a nonenveloped virus, adenovirus type 5. The inset illustrates the concept that viruses multiply by assembly of preformed components into particles. (B)Growth of an enveloped virus, Western equine encephalitis virus, a member of the Togaviridae . This virus acquires infectivity after maturation at the plasma membrane, and therefore, little intracellular virus can be detected. The small quantities observed at each time point probably represent released virus contaminating the cell extract.
Examples of global analyses include genome-wide transcriptional profiling to study the host response to infection. Introduction of the 1918 strain of influenza virus into mice leads to a rapidly fatal disease characterized by sustained induction of proinflammatory cytokine and chemokine genes. Understanding the gene expression signature that correlates with lethality is one goal of these studies. Global analysis can also predict signatures of vaccine efficacy. In one study, transcriptional profiling of peripheral blood mononuclear cells from vaccinated subjects revealed that the yellow fever virus vaccine induces the expression of genes encoding members of the complement system and stress response proteins. This pattern accurately predicts CD8 +T cell and antibody responses that are thought to mediate protection from infection with yellow fever virus. A separate signature that accurately predicts neutralizing antibody synthesis during infection was also identified.
Some of the methods used in global analysis are described below.
DNA Microarrays
An early staple of global analyses, this method enables the study of the gene expression profile of a cell in response to virus infection ( Chapter 14) and can also be used to discover new viruses. In this method, millions of unique viral DNA sequences fixed to glass or silicon wafers are incubated with sequences complementary to DNAs or RNAs, which have been amplified from clinical and environmental samples by PCR. Binding is usually detected by using fluorescent molecules incorporated into amplified nucleic acids. Microarrays have been largely supplanted by high-throughput sequencing, which allows identification of transcripts and their quantification in an unbiased manner, e.g., without prior assumption of what genes are involved.
In RNAseq, RNAs extracted from cells or tissues are converted by reverse transcription to complementary DNAs, which are then subjected to high-throughput DNA sequencing. The results provide insight into sequences and quantity of RNAs in a cell at a given time under specific conditions. It allows detection and quantification of transcripts that are not represented on microarrays. Information on transcriptional activity is provided by native elongating transcript sequencing(NET-seq), in which immunoprecipitation of RNA polymerase is followed by high-throughput sequencing of the 3′ ends of the associated RNAs. A method to study the association of RNAs with ribosomes is ribo-seq, in which polysomes are treated with RNases and the 20- to 30-nucleotide ribosome-protected fragments are sequenced. The information provides insight into translational control of gene expression and the mechanism of protein synthesis and allows annotation of translated sequences.
A number of methods yield global views of protein-nucleic acid interactions at unprecedented levels of resolution. Chromatin-immunoprecipitation sequencing( ChiP-seq) can localize protein-DNA interactions with single-nucleotide precision ( Fig. 2.20). In this method, protein-DNA complexes are immunoprecipitated with antibodies to DNA binding proteins, such as transcription proteins, histones, or even specific methyl groups on histones. The DNAs are then subjected to high-throughput sequencing to identify the sites on DNA to which these proteins bind. An early variant called ChiP on chipemployed microarrays to identify protein binding sites on DNA.
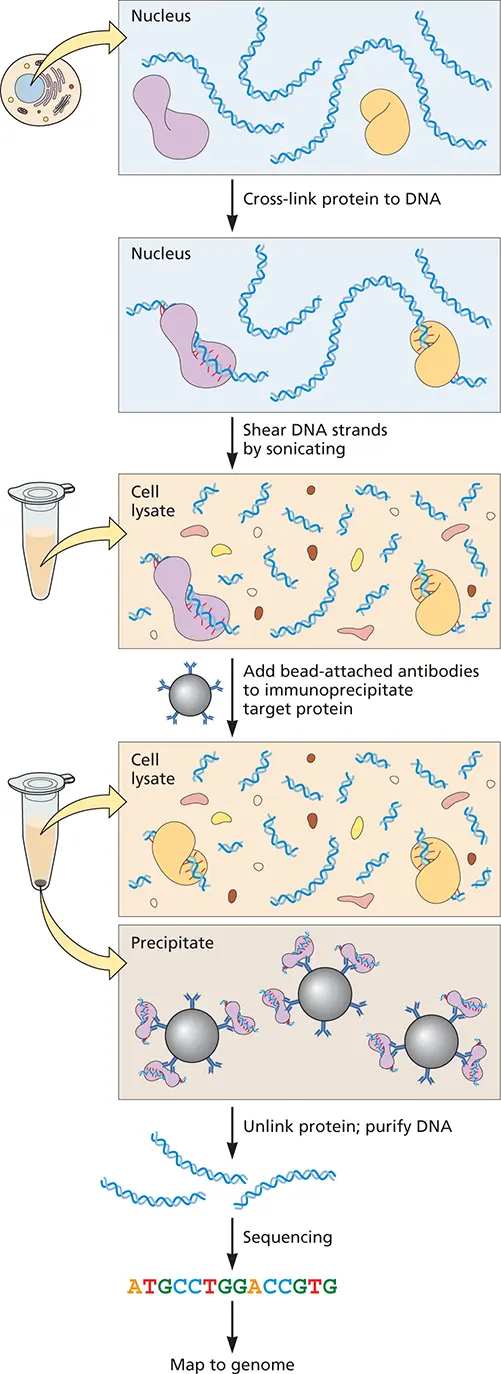
Figure 2.20 Chromatin immunoprecipitation and DNA sequencing, ChiP-seq. This technique is used to identify the precise binding sites of proteins on DNA. DNA is cross-linked to proteins by treating cells with formaldehyde, followed by sonication to shear DNA to 200 to 1,000 bp. Beads coated with antibody to the DNA binding protein of interest are added and precipitated. The protein is removed and DNA purified and subjected to high-throughput sequencing to identify protein binding sites on the DNA.
Many protocols have been devised for genome-wide analysis of RNA-protein interactions that are based on cross-linking immunoprecipitation(CLIP). In CLIP-seq, RNA-protein complexes are cross-linked in cells in culture with UV light. Cells are lysed and proteins of interest are immunoprecipitated. Proteins are removed by digestion with protease, DNA is synthesized from the previously bound RNA with reverse transcriptase, and the product is subjected to high-throughput sequence analysis. Interaction sites are identified by mapping the nucleic acid sequence reads to the transcriptome. A modification of this technique is called photoactivatable ribonucleoside-enhanced cross-linking and immunoprecipitation, PAR-CLIP. In this method, photoreactive ribonucleoside analogs such as 4-thiouridine are incorporated into RNA transcripts in living cells. Irradiation with UV light induces efficient cross-linking of RNAs containing these analogs to interacting proteins. Immunoprecipitation and sequencing are then carried out as in other CLIP methods.
Читать дальшеИнтервал:
Закладка:
Похожие книги на «Principles of Virology»
Представляем Вашему вниманию похожие книги на «Principles of Virology» списком для выбора. Мы отобрали схожую по названию и смыслу литературу в надежде предоставить читателям больше вариантов отыскать новые, интересные, ещё непрочитанные произведения.
Обсуждение, отзывы о книге «Principles of Virology» и просто собственные мнения читателей. Оставьте ваши комментарии, напишите, что Вы думаете о произведении, его смысле или главных героях. Укажите что конкретно понравилось, а что нет, и почему Вы так считаете.











