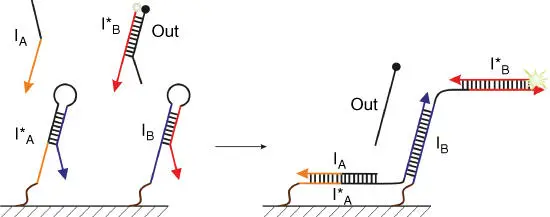
Figure 4.4DNA strand displacement logic gates attached to a DNA origami tile. Input strand I Abinds to hairpin I* Aand unwinds it. The unwound end interacts with a closely placed hairpin I B, which consequently becomes unwound and encounters a fluorophore‐labeled strand I* B, with fluorescence being attenuated by the quencher‐carrying strand Out. I Breassociates with I* B, which leads to increase in fluorescence.
The decrease in signal intensity associated with the increase in the number of integrated gates was observed for the propagation of information through the chain of the conjugated gates. This is an expected limitation of the approach, since the presence of only one oligonucleotide input provides only limited contribution to the stabilization energy for the DNA strand association (e.g. shown in Figure 4.4): additional energy input is needed to ensure robust communication of the conjugated gates. A combination of both strand displacement and deoxyribozyme logic gates [59,60] can address this issue, since an active RCDZ can be produced andused for fuelingthe cascades by cleaving RNA phosphodiester bonds.
4.4 Logic Gates Connected Via DNA Four‐Way Junction (4WJ)
Arguably, any DNA‐based sensor can be converted into a set of DNA logic gates. One of the most elegant nucleic acid sensors is the molecular beacon (MB) probe [61,62], a fluorophore‐ and a quencher‐conjugated DNA hairpin ( Figure 4.5a). MB probes have been used in DNA analysis in biomedical practice [62,65]. We designed a series of logic gates using an MB probe as a convenient fluorescent reporter [27,63,64,66,67]. Figure 4.5b,c demonstrates two designs for DNA YES gates. In both designs, DNA strands associate to form a structure stabilized through the formation of 4WJ, which gave the name to the DNA gates: “4WJ‐1” and “4WJ‐2.” 4WJ‐1 YES gate consists of two oligoethylene glycol‐modified DNA strands A and B and an MB probe. These three oligonucleotides coexist in solution in a dissociated state when a nucleic acid analyte is absent; MB is in the stem‐loop conformation, and the fluorescent signal is low. Addition of a DNA input leads to the cooperative hybridization of strands A and B both to the analyte and to the MB probe, which results in the formation of a 4WJ‐containing complex ( Figure 4.5b, bottom). The fluorophore and the quencher are remote from each other in this complex, which results in high fluorescence. Addition of oligoethylene glycol linkers to strands A and B was required to ensure the elongated conformation of the MB probe in the 4WJ complex [63]. YES gate in 4WJ‐2 design responds to the input presence according to a similar scheme. However, strands O and C constituting the gate do not require oligoethylene glycol modifications, since the two MB‐binding arms of strand C ensured the required elongated MB conformation in the input‐bound complex [64].
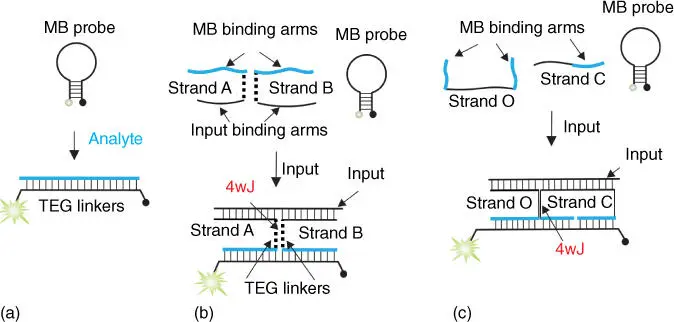
Figure 4.5Design of molecular beacon (MB)‐based DNA logic gates. (a) MB probe fluoresces upon bind complementary analyte. (b) The 4WJ‐1 design for a YES gate. Oligoethylene glycol linkers triethylene glycol (TEG) are shown as dashed lines.
Source: Based on Lake et al. [63]
. (c) The 4JW‐2 design of YES gate.
Source: Based on Cornett et al. [64].
We have designed and optimized individual 4WJ logic gates in solution with the main purpose of integrating them into computational circuits [63,64,66,67]. The designs of NOT and AND gates are presented in Figure 4.6a,b. NOT gate forms a fluorescent complex with an MB probe; addition of an oligonucleotide input disrupts the 4WJ structure, which results in the release of the MB probe from the NOT complex yielding low fluorescent signal. AND gate consists of three DNA strands folded in hairpin structures ( Figure 4.6b). Two oligonucleotide inputs open their correspondent AND hairpins and trigger association of the two strands with the MB probe. Again, the complex is stabilized by the 4WJ structure. The output of the gates is a DNA fragment formed by the portions of the two AND strands (e.g. blue and green in Figure 4.6b). Therefore, the 4WJ gate design preserves the input–output homogeneity, which is important for the integration of the gates into circuits, since an output of one 4WJ gate can be recognized as an input by the downstream gate. We achieved three layers of gate integration in solution by building an XOR logic from one OR, two AND, and two NOT gates [66]. Expectedly, the signal intensity would decrease upon adding each new level of integrated gates [66].
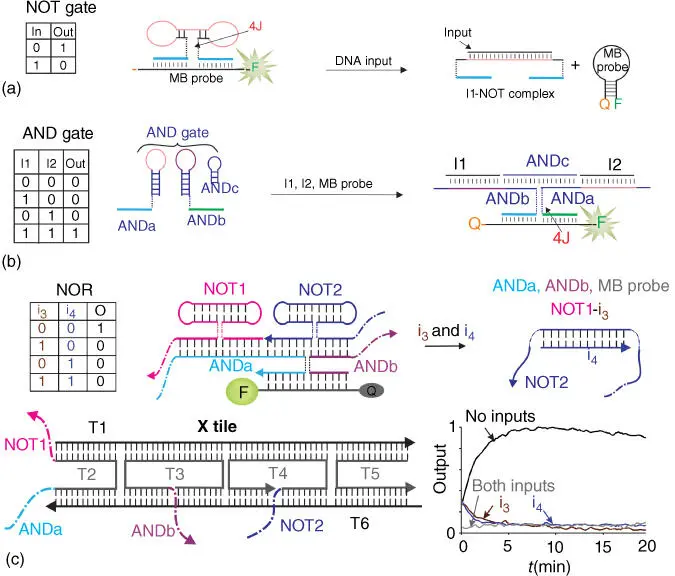
Figure 4.64WJ DNA logic gates and tile‐integrated DNA circuits. (a) 4WJ NOT gate: a DNA strand (NOT) holds the opened MB probe in the absence of an input. Addition of the DNA input decomposes the complex, thus releasing the MB probe. (b) Two‐input 4WJ AND gate. The gate consists of ANDa, ANDb, ANDc, and an MB probe. The five‐stranded 4WJ association is formed only in the presence of both inputs I1 and I2. (c) Two‐input 4WJ NOR gate integrated into a DNA tile. NOT1, NOT2, ANDa, and ANDb strands are attached to a DNA crossover (X) tile at the indicated points. In the absence of inputs, the 5′‐terminal NOT1 and 3′‐terminal NOT2 output fragments are bound to the input‐recognition fragments of the ANDa and ANDb strands, respectively, which enables formation of the high‐signal (fluorescent) 4WJ NOR association. Addition of inputs i3 and/or i4, which are complementary to the input‐recognition fragments of NOT1 and NOT2, respectively, results in dissociation of the 4WJ NOR complex. The gate performs as predicted according to the observed changes in fluorescent output (lower right corner).
To facilitate inter‐gate communication and reduce the undesirable crosstalk, we attempted to confine the gates in a nano‐environment by attaching them to a DNA scaffold in proper orientation and near their direct communication partners [27]. For example, Figure 4.6c demonstrates integration of two NOT with an AND gate to produce a NOR logic: the two NOT gates feed their outputs to a 2iAND. The NOT and AND strands form a fluorescent complex with an MB probe, while addition of at least one input decomposes one of the NOT gates followed by the collapse of the entire complex into the separate strands. The obtained circuits were shown to be reusable multiple times if RNA inputs are used in the presence of RNase H as a buffer component. As the case for strand displacement‐based logic gates, a universal powering method is needed to build long chains of communicating logic gates.
Majority of the DNA logic gates, however, explore only two to five layers of integration, which faces significant signal reduction as the signal propagates along the chain of communicating gates. At least partially, this problem can be mitigated by localizing logic gates in a specific order and at precise positions on a DNA tile for efficient communication as it is used in electronic processors. An energy input is required to “push” the signal through the DNA association, an approach that has not been realized yet. Alternatively, parallel computation using multiple small‐scale integrated circuits can be explored. While all the technical problems can be eventually addressed given the appropriate time and effort, the future of molecular computation depends on the practical usability of DNA computers. Indeed, it becomes clear that computers based on hybridization of DNA strands cannot compete with electronic devices in terms of the processing speed due to much slower rates of DNA hybridization than electron transfer in semiconductor materials. Instead, biocompatible and biodegradable DNA‐based logic constructs can be used for manipulating biological molecules and objects (cells), which can eventually find applications in addressing biological and biomedical problems.
Читать дальше















