The study of DNA nanoparticles has been one of significant interest since the early 1980s due to the progressive developmental foundation proposed by Nadrian Seeman [16,17]. Since the conceptual layout of DNA nanotechnology was proposed, widespread interest toward DNA nanotechnology [18–20] and RNA nanotechnology [21–24] has occurred in the scientific community. The design of nucleic acid nanoparticles into well‐defined two‐ or three‐dimensional shapes can be accomplished by using the DNA origami technique [25,26]. This approach utilizes designing multiple short DNA fragments (“staple” strands) that force folding of a long single‐strand DNA (DNA template) into a preprogrammed shape. Computational tools are used to calculate the placement of individual staple strands within a specific region of the DNA template, and due to Watson–Crick base pairing, the necessary sequences of all staple strands can be executed. Figure 5.1demonstrates an example of the DNA dolphin‐shaped structure obtained by DNA origami [27]. Inspired by the DNA origami technique, researchers developed RNA origami. In this approach, RNA polymerase is implemented to transcribe a long RNA strand (RNA template) that can fold into a pre‐rendered shape at isothermal conditions without a need for staple strands [29]. More often, RNA nanoparticles are designed using known crystal structures of complex RNA molecules such as ribosomal RNA containing multiple, well‐defined, and often recurrent RNA structural motifs [30]. These motifs are then manually extracted from larger RNA complexes using 3D modeling software such as Swiss‐PDBViewer [31]. RNA motifs serve as modular building blocks that can be further interconnected to obtain a desired shape [22,28,32,33]. As a result, infinite numbers of nucleic acid nanoparticles with intricate shapes and dimensions can be modeled and assembled utilizing above techniques as exemplified in Figure 5.1. Researchers have used nucleic acids, both DNA and RNA, to fabricate artificial nucleic acid complexes for a variety of applications [34–40]. This has led to the development of therapeutic nucleic acid nanotechnology [41,42], various devices for structure probing in vitro and in vivo [43,44], and biomimetic systems [45], as well as development of nucleic acid “smart” devices capable of performing simple and complex molecular computations [43,46].
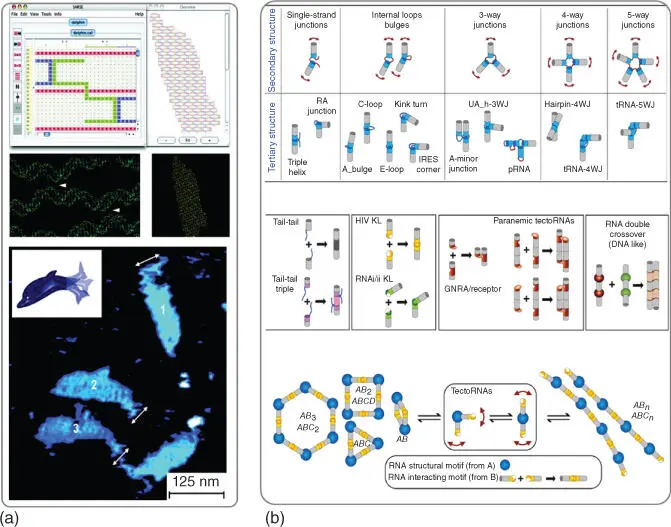
Figure 5.1Nucleic acid nanostructure designing techniques. (a) DNA origami method was used to computationally design and assemble a dolphin‐shaped DNA nanostructure. (b) Example of RNA nanoparticle design approach. Variety of RNA tertiary structures (tecto‐RNAs) were combined to construct different nano‐objects of 1D or linear shape, 2D or polygonal shapes, and 3D shapes.
Source: (Panel a) From Andersen et al. [27]. Reprinted with the permission of American Chemical Society; (Panel b) From Grabow and Jaeger [28]. Reproduced with the permission of American Chemical Society.
5.2 RNA Aptamers are Modular and Programmable Biosensing Units
In parallel to the nucleic acid nanotechnology field, scientific attention has been drawn to nucleic acid aptamer development and their implementations. A nucleic acid aptamer consists of RNA and/or DNA oligonucleotides that bind to a specific target moiety (ligand) with high affinity and specificity. A wide range of various chemical and biological entities can serve as ligands ranging from a small molecule to a mammalian cell [47]. The development of aptamer oligonucleotides that bind to specific ligands was engineered through repeated rounds of in vitro selection also known as SELEX (systematic evolution of ligands by exponential enrichment) in the early 1990s in two independent groups of Larry Gold and Jack Szostak [48,49]. Since then, numerous different types of aptamers were engineered in academic research labs and found applications in various biotechnological fields ( Figure 5.2a). Several recent reviews provide a critical evaluation of nucleic acid aptamer technology with their applications in vitro and in vivo [50–54].
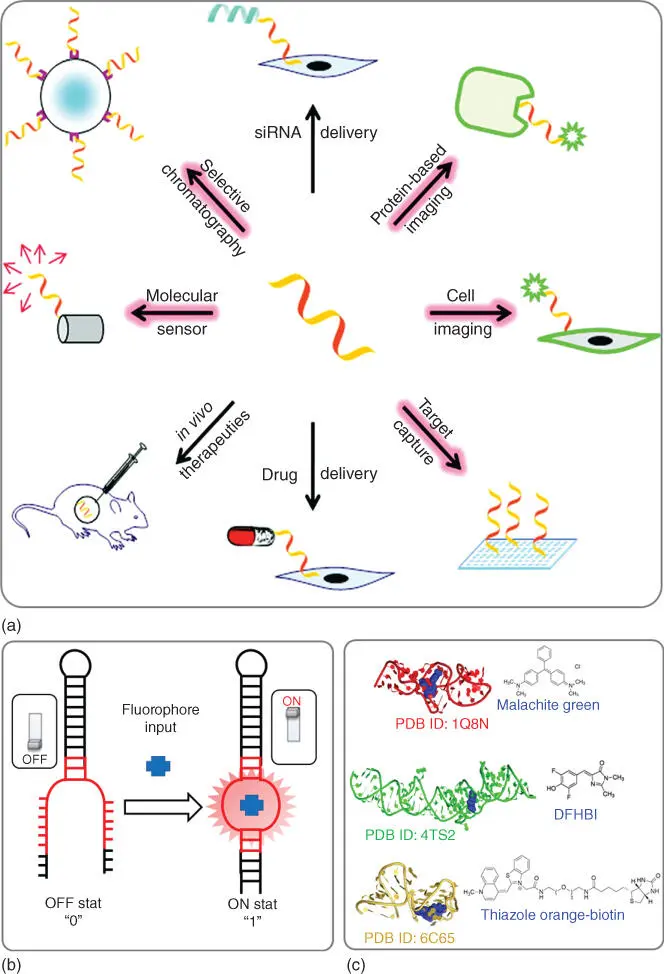
Figure 5.2(a) Examples of the diverse application of nucleic acid aptamers. (b) Schematic 2D representation of fluorogenic RNA aptamer YES gated function. (c) Examples of known light‐up RNA 3D structures of MG‐binding aptamer (red), Spinach RNA aptamer (green), and Mango RNA aptamer (gold) with corresponding fluorophore ligands.
Source: (Panel a) From Iliuk et al. [50]. Reproduced with the permission of American Chemical Society.
In this chapter, we are primarily focusing on fluorogenic RNA aptamers, also known as light‐up RNA aptamers. In particular, discussions will cover fluorogenic RNA aptamers as potential candidates to be used in biocomputing applications. Utilizing light‐up aptamers in bioanalytical disciplines for an analyte sensing in vivo has several advantages. For instance, RNA aptamers can be genetically fused to genes coding for cellular RNAs of interest that in turn can be advantageous in endogenous synthesis of the targeting sequence [55–58]. Inserting light‐up aptamers directly into the target endogenous RNA can be restricted by the labor‐intensive genetics. However, these aptamers, even when used in multiple copies, do not significantly increase the size of the cellular RNA. Only the imaging dye needs to be introduced exogenously, and usually small nonpolar organic dyes can be passively diffused into cells. The major concerns of applying unmodified RNA aptamers in vivo are their susceptibility to degradation by nucleases, making it difficult to intracellularly express them. To tackle this problem, aptamer molecules are expressed by being inserted into a complex noncoding RNA strand, like tRNA, or into an RNA junction to protect the aptamer from degradation and induce folding. Such insertion of the RNA aptamer sequence into endogenous RNA not only allows transcription to be monitored in vitro but can also be used to regulate gene expression in a logical manner [59,60].
In a fluorogenic RNA aptamer, the ligand is typically a small organic dye that can emit fluorescent light upon binding to its aptamer host molecule (otherwise, it is in a nonfluorescent state). This feature is very attractive to develop binary ON and OFF systems where a fluorescence state is ON and a nonfluorescence state is OFF as shown in Figure 5.2b. Over the past two decades, several RNA fluorogenic aptamers were developed, and all can be potentially implemented for fabrication of a nanocomplex with a simple YES logic operation with fluorescence outputs ON and OFF. The examples of such RNA aptamers and ligands as well as their properties are summarized in Table 5.1, and structures of the three most common RNA aptamers are illustrated in Figure 5.2c. The interaction between a ligand and a binding pocket of the host aptamer is usually non‐covalent. Therefore, the interaction can be easily reversed by introducing denaturing agents or certain conditions that will disrupt the correct pocket conformation. Such disruption prevents ligand binding and thus serves as input signals. Once the ligands are in an unbound state (free in solution), their intrinsic fluorescence emission is diminished.
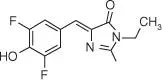
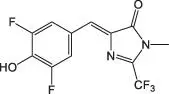
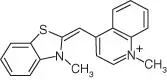
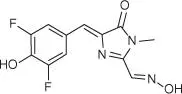
Table 5.1List of some common RNA aptamers with corresponding fluorogenic ligands and properties.
| Fluorogen dye |
Light‐up aptamer |
K D(nM) |
Ex/Em (nm) |
ℰ (M −1/cm) |
Φ a) |
Fluorogen structure |
PDB ID b) |
References |
| OTB |
DiR2s‐Apt |
662 |
380/421 |
73 000 |
51 |
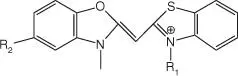 |
6DB9 |
[61] |
| DFHBI |
Spinach |
540 |
469/501 |
24 300 |
72 |
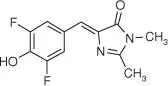 |
4TS0 |
[62] |
| DFHBI‐1T |
Spinach2 |
560 |
482/505 |
31 000 |
94 |
 |
6B3K |
[63] |
| DFHBI‐2T |
Spinach2 |
1300 |
500/523 |
29 000 |
12 |
 |
6B3K |
[63] |
| TO‐1 |
Mango |
3 |
510/535 |
77 500 |
14 |
 |
5V3F |
[64] |
| DFHO |
Corn |
70 |
505/545 |
29 000 |
25 |
 |
6E80 |
[65] |
| DIR |
DIR apt |
86 |
600/646 |
134 000 |
26 |
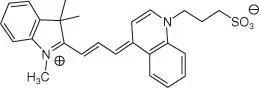 |
3T0W |
[66] |
| Mal. Green |
MG aptamer |
117 |
630/650 |
150 000 |
19 |
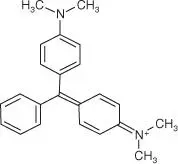 |
1Q8N |
[67] |
| DIR‐pro |
DIR2s‐Apt |
252 |
600/658 |
164 000 |
33 |
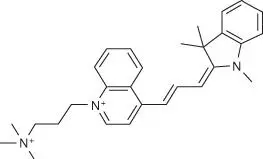 |
6D89 |
[61] |
| TO‐3 |
Mango |
6–8 |
637/658 |
9300 |
N/A |
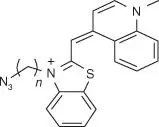 |
5V3F |
[64] |
aΦ referred to quantum yield of the complex expressed in percentage.
Читать дальше
























