bProtein Data Bank ID number.
A highly effective RNA‐based fluorogenic unit should possess specific features. The ideal dye needs to display a high absorption coefficient ( ɛ ) to ensure sensitive detection and to minimize fluorescence background. The fluorophore should show a low ratio of photons absorbed to photons emitted (quantum yield), meaning it should have a high fluorescence enhancement and brightness. The RNA–fluorophore interaction should be highly specific and occur with high affinity to make it possible to use low concentrations while still obtaining high contrast and keeping background fluorescence low. The aptamer–fluorophore complex also needs to be photostable to extend the ability for data acquisition. Often, these types of fluorogenic aptamers have superior characteristics over the electrochemical and colorimetric approach for sensing and imaging. For example, RNA strands do not require chemical conjugation, and the RNA aptamer provides high sensitivity and high speed of response while also exhibiting high spatial resolution [68,69].
Malachite green (MG)‐binding RNA aptamer was one of the earliest models of RNA light‐up aptamer and was extensively studied in laboratory settings [38–40,43,46,67,70,71]. Excitation of free triphenylmethane fluorophore (MG) in a solution results in low fluorescence due to easy vibrational de‐excitation (i.e. excess energy from the MG excited state is dissipated in the form of vibrational movement). When MG is bound to its RNA aptamer, MG is stabilized in a planar form, and vibrations are restricted, which results in a 2000‐fold increase of fluorescence [67]. In nucleic acid nanofabrication, fluorescent aptamers can potentially act as fluorescent reporter units that can be harbored into a larger complex nanoparticle by simple extension of individual strands. Several previous reports have shown that MG‐binding aptamers can be used for co‐transcriptional assembly verification [43] as well as monitoring of the dynamic behavior of interdependent RNA–DNA hybrids [35].
More recently, a novel and much less intracellular toxic RNA aptamer, as compared with MG RNA aptamer, Spinach RNA aptamer, was developed [62]. The Spinach RNA aptamer binds the green fluorescent protein (GFP) fluorophore analog DFHBI (( Z )‐4‐(3,5‐difluoro‐4‐hydroxybenzylidene)‐1,2‐dimethyl‐1 H ‐imidazol‐5(4 H )‐one) [72]. The work on the Spinach aptamer has been extended to produce Spinach 2 that has much greater thermostability and brightness. However, Spinach 2 is more susceptible to degradation by nucleases [63,73]. Further effort has been made to develop yet another “vegetable” aptamer called Broccoli that binds DFHBI‐1T (derivative of the DFHBI dye) [74].
The interaction between the aptamer and its target often causes slight structural rearrangement in favor of stabilization of the RNA–ligand complex. This feature can be used to control RNA–ligand binding allosterically, where the allosteric site (sensing module) can be connected to the aptamer region (reporting module) through a communication module. This strategy was developed by Ronald Breaker using ribozymes as reporting modules [75,76], and later a similar strategy was implemented using MG‐binding RNA aptamer as a reporter unit [77,78]. Allosteric biosensors can also be used for protein detection for specific applications [79]. With these biosensors, target metabolite molecules as well as enzymes participating in an intracellular pathway can be identified. For instance, RNA biosensors are now commonly used to sense the presence of the following metabolites: cyclic AMP [80], cyclic di‐AMP [73], S ‐adenosylmethionine (SAM) [81], FMN [78], S ‐adenosyl‐L‐homocysteine (SAH) [82], and thiamine pyrophosphate (TPP) [83]. Also, the development of these aptamers has simplified RNA imaging in mammalian, yeast, and bacterial cells [44,84–87].
5.3 Construction of RNA Nanoparticles with Integrated Logic Gate Operations Using Light‐Up Aptamers
This section is focused exclusively on the fluorogenic RNA aptamer‐based methods to fabricate responsive logic‐gated nanodevices. Based on the aforementioned properties, RNA‐based light‐up aptamers are ideal for binary logic system development. These systems can be programmed to respond to an input signal, inducing conformational change within the RNA aptamer's binding pocket that will further dictate binding strength of a fluorescent dye molecule. Various factors can serve as inputs. Nonspecific factors include temperature, pH, and ionic strength. Specific factors can include predesigned competitive short oligonucleotide strands, nonfluorescent molecule mimicking structures of the ligand dye, and RNA‐binding protein biomolecules, to name a few. The concentration of the inputs often needs to be fine‐tuned to display desirable outcomes and define the threshold between ON and OFF values. Short DNA oligonucleotides are commonly used as inputs because they are relatively inexpensive, are stable in aqueous solutions, and can hybridize with DNA as well as RNA strands to form RNA–DNA duplexes. When the inputs are present at a certain concentration, the overall fluorogenic aptamer structure can mimic a computer's function toggling between fluorescence (ON) and nonfluorescence (OFF) states. However, these can be achieved only when specific LOGICAL conditions are satisfied following Boolean algebra or function. The Boolean algebra is used to analyze and simplify the digital (logic) circuits and uses only the binary numbers 0 and 1. Other values include “YES and NO” or “ON and OFF,” and this type of notation is often referred to as a binary algebra. Logic gate plays a role as an elementary building block of digital circuits. Depending on the complexity of the operations and tasks, some circuits may have only a few logic gates, while others, such as microprocessors, have combinatorial circuits embedding millions of logic gates.
5.3.1 Implementation of MG‐Binding RNA Aptamer to Design Binary Logic Gates
The biochemical applications of logic gates include biosensing and data processing [88]. The simplest logic gates are based on one input and correspond with YES and NOT gates. All fluorogenic (light‐up) RNA aptamers are macromolecules possessing YES logic functions since they turn ON only in the presence of input fluorophore molecules ( Figure 5.2b). Examples of other common logic gates requiring two inputs implemented in binary algebra and their truth tables defining each function are summarized in Figure 5.3. The six most common binary logic gates include AND, OR, XOR, NAND, NOR, and XNOR. Among these gates, the OR logic gate yields 1 output when at least one of the inputs equals 1. The AND logic gate generates 1 output if and only if both inputs are 1. It is important to note that theoretically to form a complete set of all possible logic gates, only four basic YES, NOT, AND, and OR gates are required.
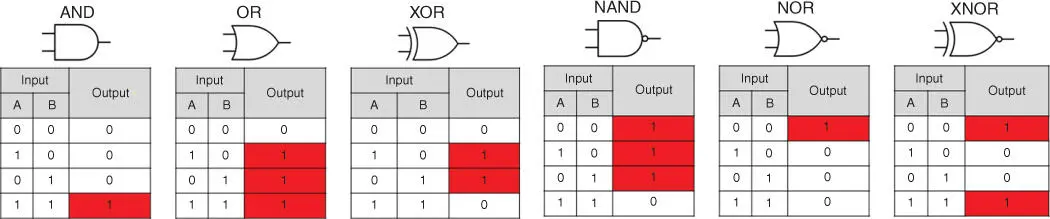
Figure 5.3Common binary logic gate symbols and truth tables.
The RNA light‐up aptamers are the perfect system to develop devices operating in a gated manner. However, currently there only a handful of reports assessing fluorogenic aptamers from this perspective [46,89–92]. Recent work reported by Goldsworthy et al. [46] notably utilized MG‐binding RNA aptamers to design complex systems performing AND, OR, NAND, and NOR logic gate functions. The design principle is illustrated in Figure 5.4a. In AND and OR specific examples, the MG‐binding RNA aptamer sequence (shown in representative red color) was extended at each 5′‐ and 3′‐ends to have 26 nts programmed to interfere with the structure of the MG‐binding pocket. The resulting AND and OR gates with default setting 0‐0 (no inputs present) produced 0 output. Two short DNA oligonucleotides served as inputs (A and B) that hybridize with the interfering ends (shown in Figure 5.4in green and blue colors). The AND logic achieved output 1 (ON conformation) only when two inputs were present at the same time. The OR logic function is converted into the ON state when at least one of the inputs was present. Since the RNA sequences for the AND and OR operations were the same, inputs of varying length were chosen to selectively manipulate the response from the MG‐binding RNA aptamer.
Читать дальше













