Jane Flint - Principles of Virology
Здесь есть возможность читать онлайн «Jane Flint - Principles of Virology» — ознакомительный отрывок электронной книги совершенно бесплатно, а после прочтения отрывка купить полную версию. В некоторых случаях можно слушать аудио, скачать через торрент в формате fb2 и присутствует краткое содержание. Жанр: unrecognised, на английском языке. Описание произведения, (предисловие) а так же отзывы посетителей доступны на портале библиотеки ЛибКат.
- Название:Principles of Virology
- Автор:
- Жанр:
- Год:неизвестен
- ISBN:нет данных
- Рейтинг книги:3 / 5. Голосов: 1
-
Избранное:Добавить в избранное
- Отзывы:
-
Ваша оценка:
- 60
- 1
- 2
- 3
- 4
- 5
Principles of Virology: краткое содержание, описание и аннотация
Предлагаем к чтению аннотацию, описание, краткое содержание или предисловие (зависит от того, что написал сам автор книги «Principles of Virology»). Если вы не нашли необходимую информацию о книге — напишите в комментариях, мы постараемся отыскать её.
Volume I: Molecular Biology
Volume II: Pathogenesis and Control
Principles of Virology, Fifth Edition
Principles of Virology — читать онлайн ознакомительный отрывок
Ниже представлен текст книги, разбитый по страницам. Система сохранения места последней прочитанной страницы, позволяет с удобством читать онлайн бесплатно книгу «Principles of Virology», без необходимости каждый раз заново искать на чём Вы остановились. Поставьте закладку, и сможете в любой момент перейти на страницу, на которой закончили чтение.
Интервал:
Закладка:
Each monoclonal antibody binds specifically to 8 to 12 residues that fit into the antibody-combining site. These amino acids are either next to one another either in primary sequence ( linear epitope) or in the folded structure of the native protein ( nonlinearor conformational epitope). In contrast, polyclonal antibodiescomprise the repertoire produced in an animal against the many epitopes of an antigen. Antigenic sites may be identified by cross-linking a monoclonal antibody to the virus and determining which protein is the target of that antibody. Epitope mapping may also be performed by assessing the abilities of monoclonal antibodies to bind synthetic peptides representing viral protein sequences. When the monoclonal antibody recognizes a linear epitope, it may react with the protein in immunoblot analysis, facilitating direct identification of the viral protein harboring the antigenic site.
An elegant understanding of antigenic structures has come from the isolation and study of variant viruses that are resistant to neutralization with specific monoclonal antibodies (called monoclonal antibody-resistant variants). By identifying the amino acid change(s) responsible for this phenotype, the antibody-binding site can be located and, together with three-dimensional structural data, can provide detailed information on the nature of antigenic sites that are recognized by neutralizing antibodies (see the figure).
Hemagglutination inhibition.Antibodies against viral proteins with hemagglutination activity can block the ability of virus to bind red blood cells. In this assay, dilutions of antibodies are incubated with virus, and erythrocytes are added as outlined above. After incubation, the titer is read as the highest dilution of antibody that inhibits hemagglutination. This test is sensitive, simple, inexpensive, and rapid, and can be used to detect antibodies to viral hemagglutinin in animal and human sera. For example, hemagglutination inhibition assays were used to identify individuals who had been infected with the newly discovered avian influenza A (H7N9) virus in China during the 2013 outbreak.
Visualization of proteins.Antibodies can be used to visualize viral or cellular proteins in infected cells or tissues. In direct immunostaining, an antibody that recognizes a viral protein is coupled directly to an indicator such as a fluorescent dye or an enzyme ( Fig. 2.12). A more sensitive approach is indirect immunostaining, in which a second antibody is coupled to the indicator. The second antibody recognizes a common region on the virus-specific antibody.
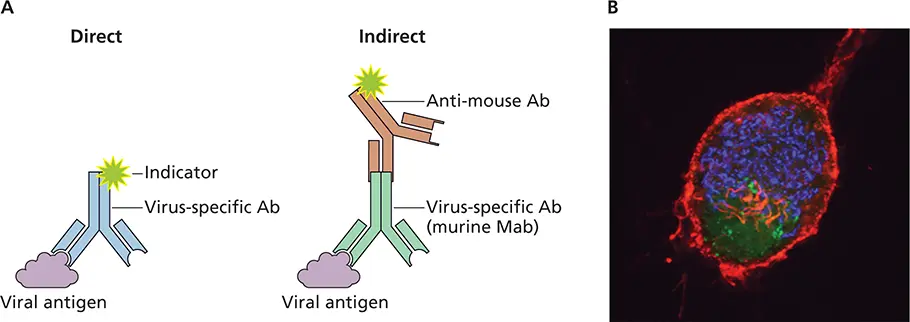
Figure 2.12 Direct and indirect methods for antigen detection. (A)The sample (tissue section, smear, or bound to a solid phase) is incubated with a virus-specific antibody (Ab). In direct immunostaining, the antibody is linked to an indicator such as fluorescein. In indirect immunostaining, a polyclonal antibody, which recognizes several epitopes on the virus-specific antibody, is coupled to the indicator. Mab, monoclonal antibody. (B)Use of immunofluorescence to visualize pseudorabies virus replication in neurons. Superior cervical ganglion neurons were grown in culture and infected with a recombinant virus that produces green fluorescent protein (GFP) fused to the VP26 capsid protein. Neurons were stained with AF568-phalloidin, which stains actin red, and anti-GM130 to stain the Golgi blue. GFP-VP26 is visualized by direct fluorescence. Courtesy of L. Enquist, Princeton University.
Multiple second-antibody molecules bind to the first antibody, resulting in an increased signal from the indicator compared with that obtained with direct immunostaining. Furthermore, a single indicator-coupled second antibody can be used in many assays, avoiding the need to purify and couple an indicator to multiple first antibodies.
In practice, virus-infected cells (unfixed or fixed with acetone, methanol, or paraformaldehyde) are incubated with polyclonal or monoclonal antibodies ( Box 2.7) directed against viral antigen. Excess antibody is washed away, and in direct immunostaining, cells are examined by microscopy. For indirect immunostaining, the second antibody is added before examination of the cells by microscopy. Commonly used indicators fluoresce on exposure to UV light. Filters are placed between the specimen and the eyepiece to remove blue and UV light so that the field is dark, except for cells to which the antibody has bound, which emit light of distinct colors ( Fig. 2.12). Today’s optics are much better at keeping the wavelengths separated, permitting the use of different colors to detect various components in the same specimen. Antibodies can also be coupled to molecules other than fluorescent indicators, including enzymes such as alkaline phosphatase, horseradish peroxidase, and β-galactosidase, a bacterial enzyme that in a test system converts the chromogenic substrate X-Gal (5-bromo-4-chloro-3-indolyl-β-D-galactopyranoside) to a blue product. In these instances, excess antibody is washed away, a suitable chromogenic substrate is added, and the presence of the indicator antibody is revealed by the development of a color that can be visualized.
Immunostaining has been applied widely in the research laboratory for determining the sub-cellular localization of cellular and viral proteins ( Fig. 2.12), monitoring the synthesis of viral proteins, determining the effects of mutation on protein production, localizing the sites of viral genome replication in animal hosts, and determining the effect of infection on structure of the tissue. It is the basis of the fluorescent-focus assay.
Immunostaining of viral antigens in smears of clinical specimens may be used to diagnose viral infections. For example, direct and indirect immunofluorescence assays with nasal swabs or washes can detect a variety of viruses, including influenza virus and measles virus. Viral proteins or nucleic acids may also be detected in infected animals by immunohistochemistry. In this procedure, tissues are embedded in a solid medium such as paraffin, and thin slices are produced using a microtome. Viral antigens can be detected within the cells in the sections by direct and indirect immunofluorescence assays.
Enzyme immunoassay.Detection of viral antigens or antiviral antibodies can be accomplished by solid-phase methods, in which an antiviral antibody or protein is adsorbed to a plastic surface ( Fig. 2.13A). To detect antibodies to viruses, viral protein is first linked to the plastic support, and then the specimen is added ( Fig. 2.13B). Like other detection methods, enzyme immunoassays are used in both experimental and diagnostic virology. In the clinical laboratory, enzyme immunoassays are used to detect a variety of viruses, including rotavirus, herpes simplex virus, and human immunodeficiency viruses. A modification of the enzyme immunoassay is the lateral fow immunochromatographic assay, which has been used in rapid antigen detection test kits ( Fig. 2.14). The lateral fow immunochromatographic assay does not require instrumentation and can be read in 5 to 20 min in a physician’s office or in the field. Commercial rapid antigen detection assays are currently available for influenza virus, respiratory syncytial virus, and rotavirus.
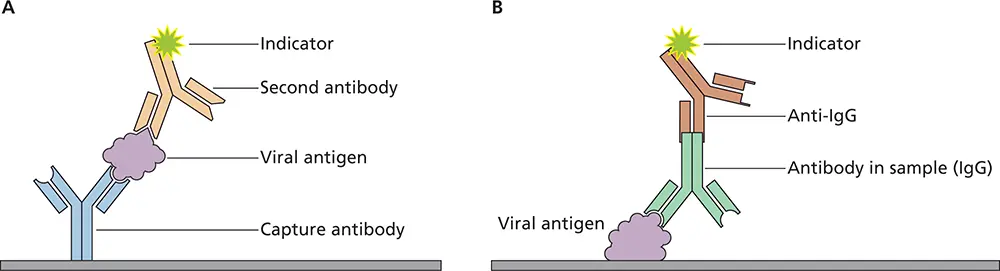
Figure 2.13 Detection of viral antigen or antibodies against viruses by enzyme-linked immunosorbent assay (ELISA). (A)To detect viral proteins in serum or clinical samples, antibodies specific for the virus are immobilized on a solid support such as a plastic well. The sample is placed in the well, and viral proteins are “captured” by the immobilized antibody. After washing to remove unbound proteins, a second antibody against the virus is added, which is linked to an indicator. The second antibody will bind if viral antigen has been captured by the first antibody. Unbound second antibody is removed by another washing, and when the indicator is an enzyme, a chromogenic molecule that is converted by the enzyme to an easily detectable product is then added. The enzyme amplifies the signal because a single catalytic enzyme molecule can generate many product molecules. Another wash is done to remove unbound second antibody. If viral antigen has been captured by the first antibody, the second antibody will bind and the complex will be detected by the indicator. (B)To detect antibodies to a virus in a sample, viral antigen is immobilized on a solid support such as a plastic well. The test sample is placed in the well, and antiviral IgG antibodies present in the sample will bind the immobilized antigen. After washing to remove unbound components in the sample, a second antibody, directed against a general epitope on the first antibody, is added. Unbound second antibody is removed by another wash. If antibodies against the virus are present in the specimen, the second antibody will bind to them and the complex will be detected via the indicator attached to the second antibody, as described in (A).
Читать дальшеИнтервал:
Закладка:
Похожие книги на «Principles of Virology»
Представляем Вашему вниманию похожие книги на «Principles of Virology» списком для выбора. Мы отобрали схожую по названию и смыслу литературу в надежде предоставить читателям больше вариантов отыскать новые, интересные, ещё непрочитанные произведения.
Обсуждение, отзывы о книге «Principles of Virology» и просто собственные мнения читателей. Оставьте ваши комментарии, напишите, что Вы думаете о произведении, его смысле или главных героях. Укажите что конкретно понравилось, а что нет, и почему Вы так считаете.











