Source: From Watson and Crick [45]. Also adapted from Zephyris, DNA Structure, Wikimedia commons, 2011. Public Domain.

Figure 1.5Leonard Adleman – a pioneer of the biomolecular computing; the photo of 1993 when the first experiments on DNA computing were running.
Source: Courtesy of Prof. Leonard Adleman.
The “traveling salesman problem” asks the following question [57–59]: “Given a list of cities and the distances between each pair of cities, what is the shortest possible route that visits each city and returns to the origin city?” It is a problem in combinatorial optimization, important in theoretical computer science. It is frequently used to test computational algorithms and computer hardware. In general, the traveling salesman problem is hard to solve, particularly when the number of the visited cities is increasing. Adleman solved the problem for seven cities only ( Figure 1.6), which was rather a trivial task, but importantly it was solved using computational power of DNA reactions [28,56]. The DNA molecules hybridized in a special way to solve the problem, and the computation was performed by numerous DNA sequences (actually rather short oligonucleotides) operating in parallel. This was important conceptual difference from Si‐based electronic computers that perform all operations in a sequence. The computation in Adleman's experiment was performed at the speed of 1014 operations per second, a rate of 100 teraflops or 100 trillion floating point operations per second (comparable to the fastest presently available quantum computer) – all because of massively parallel processing capabilities of the DNA computing operation [54,60]. The promise for extremely fast computation ignited the interest to the DNA computing concept, then being extended to a broader area of molecular [8] and biomolecular [12] computing. Despite the fact that the practical results have not been obtained after almost 25 years of the active research, optimistic expectations for building DNA computers are still present [27,55,61,62]. The advantages of the DNA and RNA computing systems are not only in their potentially high speed of operation due to the parallel information processing but also in their ability to operate in a biological environment for solving biomedical problems in terms of diagnostics and possibly therapeutic action (theranostics) [16,63], for example, for logic control of gene expression [64]. RNA‐based computing systems are particularly promising for in vivo operation, thus being excellent candidates for nanomedicine with implemented Boolean logic [65]. DNA computers can operate as a Turing machine [51] and can be sophisticated enough to mimic neural network computations similar to human brain, obviously in a very simplified way [66]. The DNA computing systems playing a tic‐tac‐toe game against human have been “smart” enough to win [13,67–69] ( Figure 1.7).
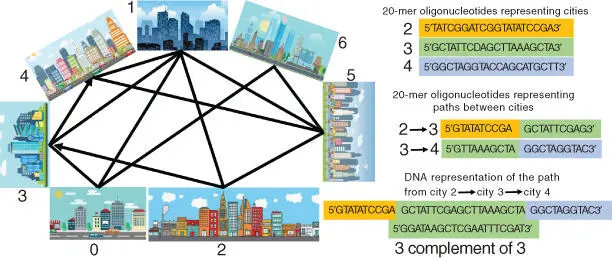
Figure 1.6The principle of Leonard Adleman's DNA computer to solve the “traveling salesman problem” (see detailed explanation in Ref. [54]).
Source: Based on Parker [54].
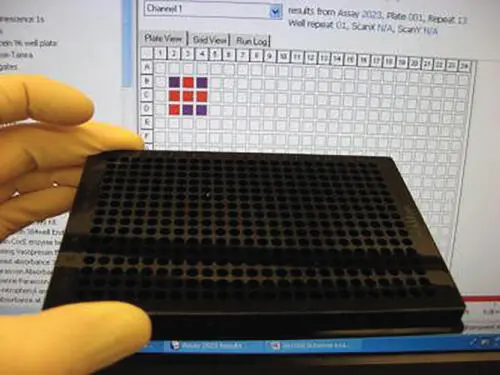

Figure 1.7The DNA computer playing the tic‐tac‐toe game. Shown in the foreground is a cell culture plate containing pieces of DNA that code for possible “moves.” A display screen (background) shows that the computer (red squares) has won the game against a human opponent (blue).
Source: Courtesy of Prof. Milan Stojanovic, Columbia University.
The progress in the DNA computing has been based on three major developments: (i) the use of sophisticated DNA structures (e.g., origami), (ii) the use of more powerful instrumentation for automatic operation of DNA computing steps (DNA chips), and (iii) specialized programming languages specifically developed for the DNA computing. The invention of the DNA origami structures [70,71] – nanoscale folding of DNA resulting in nonarbitrary two‐ and three‐dimensional shapes [72,73] ( Figure 1.8) – resulted in further sophistication of the DNA computing systems [74], capable of operating as nanorobots in living organisms [75–77]. The use of DNA microarrays (DNA chips [78]) allowed simultaneous analysis of large numbers of DNA probes [53], thus introducing a powerful hardware for the DNA computing ( Figure 1.9). A special computational language, DNA strand displacement (DSD) tool, similar to programming languages used in electronic computers, has been developed by scientists at Microsoft Research for programing DNA computing [79,80] ( Figure 1.10). The language uses DSD as the main computational mechanism, which allows devices to be designed solely in terms of nucleic acids. DSD is a first step toward the development of design and analysis tools for DSD and complements the emergence of novel implementation strategies for DNA computing. The DNA computation can be performed in living cells by DNA‐encoded circuits that process sensory information and control biological functions. A special computing language, “Cello,” has been developed for programing DNA logic operations in vivo [81]. Overall, the use of computing languages simplified the design of DNA computing systems of high complexity.
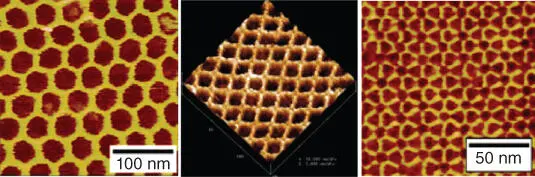

Figure 1.8Atomic force microscopy (AFM) images of DNA origami with different shapes.
Source: From Hong et al. [73]. Reprinted with the permission of American Chemical Society.
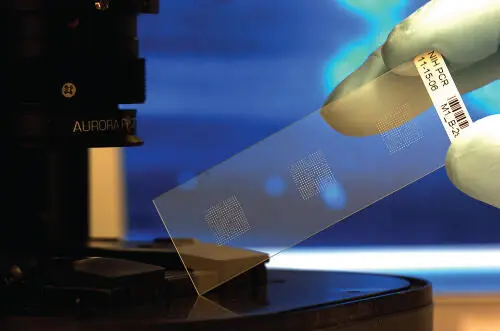

Figure 1.9An example of a DNA chip used in the DNA sensing and computing. The chip represents a DNA microarray as a collection of microscopic DNA spots attached to a solid surface. Each DNA spot contains picomoles of a specific DNA sequence. The chip allows simultaneous analysis of many DNA probes. The analysis of the probes can be performed optically (as it is in the present example) or electrochemically (then the chip should be based on a microelectrode array).
Source: Courtesy of Argonne National Laboratory and Mr. Calvin Chimes.
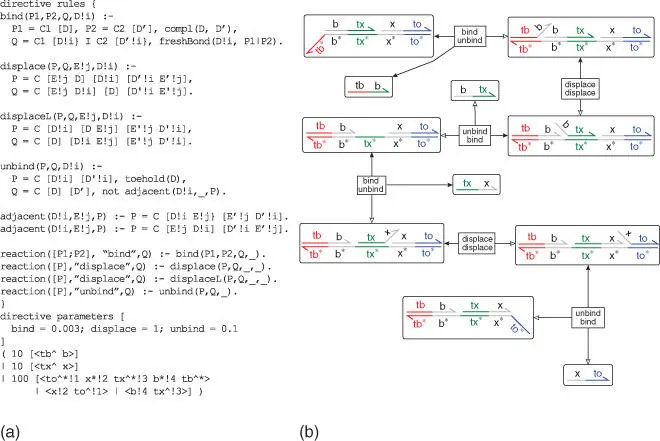
Figure 1.10Logic program (a) and automatically generated chemical reaction network (b) for a DNA strand displacement example.
Source: Adapted from Spaccasassi et al. 2019 [80] with permission; open access paper.
1.3 DNA‐Based Information Storage Systems
Human civilization generates hugeamount of information increasing exponentially and required to be stored. The total digital information today amounts to 3.52 × 10 22bits globally and at its consistent exponential rate of growth is expected to reach 3 × 10 24bits by 2040 [82]. Data storage density of silicon chips is limited, and magnetic tapes used to maintain large‐scale permanent archives begin to deteriorate within 20 years. Alternative methods/materials for storing high density/large amount of information with reliable preservation over long period of time are urgently needed. DNA has been recognized as a promising natural medium for information storage [83]. Indeed, the DNA molecules were created by nature to keep the genetic code, which can be easily “written” and “read” by biomolecular systems. With information retention times that range from thousands to millions of years, volumetric density 10 3times greater than flash memory, and energy of operation 10 8times less, DNA is a memory storage material viable and compelling alternative to electronic memory. Recent research in the area of information storage with DNA molecules resulted in the proof‐of‐the‐concept systems [82,84–87], while the practical use of the DNA memory systems is only limited by technological problems. Both processes in the information storage with DNA, “writing” and “reading” information, are available, but they are not as simple as needed to be implemented with the present computer technology. In other words, the DNA memory is technically possible, but it is not convenient enough to be integrated with standard Si‐based computers operated by end users.
Читать дальше


















