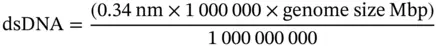2.3 Genome Sizes in the Tree of Life
There is no direct correlation between the genome size of a species and the complexity of its phenotype. In any case, the intellectual curiosity regarding the size of genomes still remains. Determination of genome size based on DNA sequencing data is one of the most accurate methods to date. To observe the lack of correlation between genome size and phenotype, upper-bound extremes can be considered here. As expected in an intuitive manner, eukaryotes show the largest genomes. In animals, the amphibian Ambystoma mexicanum (the Mexican Axolotl) shows the largest (sequenced) genome observed in nature to date. A. mexicanum shows a genome size of 32 396 Mbp (32 Gb) and a physical length that can reach up to 30 cm [166]. In plants, the record is held by Pinus lambertiana (27 603 Mbp) and Sequoia sempervirens (26 537 Mbp). P. lambertiana is the tallest and most massive pine tree [167, 168]. S. sempervirens species includes the tallest living trees on Earth (115.5 m in height or 379 ft) [169]. Among the prokaryotes, Minicystis rosea and Sorangium cellulosum So0157-2 show the largest genomes. The bacterial genome of M. rosea contains 16 Mbp of DNA (GC%: 69.1) and shows the maximum genome size found in prokaryotes [170]. Secondary to this species is the bacterial genome of S. cellulosum So0157-2 , with 14.78 Mbp of DNA (GC%: 72.1) [171]. As discussed in the previous chapter, endosymbiosis challenges the notion of the smallest genome necessary for life. The smallest prokaryotic genomes were found in different obligate symbionts. One such case is Nasuia deltocephalinicola with a genome of 112 kbp (0.11 Mbp) [172, 173]. The eukaryotes with the smallest nuclear genome necessary for life are found in the kingdom of fungi. The spore-forming unicellular parasite Encephalitozoon intestinalis shows a genome size of ∼2.3 Mbp and a total of 1.8k protein-coding genes [174]. Nonetheless, the smallest free-living eukaryote is Ostreococcus tauri , a marine green alga with a diameter of about 0.8 μm and a genome size of 12.6 Mbp (8.2k protein-coding genes) [175].
2.3.1 Alternative Methods
The data mentioned above were determined by DNA sequencing approaches made so far. DNA sequencing is an ongoing process for several decades and the species chosen for sequencing are usually either of economic or research importance (or even of historical significance). There are many species that have not yet been sequenced, either due to their minor importance to humans or due to large genomes that cannot be easily managed. Usually, the size of the genetic material can be estimated by methods other than sequencing. One of these methods is flow cytometry, which estimates the weight of the genetic material [176]. This weight, expressed in picograms (pg), can then be converted to base pairs. One picogram is equal to 978 megabase pairs (1 pg = 978 Mbp) [177]. For instance, Paris japonica (flower) shows a genome weight of 152.23 pg, which suggests a genome size of 148 880 Mbp (152.23 pg × 978 Mbp = 149 Gbp) [178].
2.3.2 The Weaving of Scales
To get a sense of genome size closer to our reference system, some transformations can express the mega base pairs as physical lengths. The linear length of a double-stranded DNA (dsDNA) molecule can be calculated by multiplying the average distance between bases (∼3.4 angstrom = 0.34 nm [179, 180]; 1 angstrom = 0.1 nm) by the total number of base pairs in a genome. Here, genomes are expressed in mega base pairs. Since 1Mbp is equal to one million base pairs, the size of a genome can be multiplied by one million and then multiplied further by the average distance between bases (0.34 nm). One meter is equal to 1 000 000 000 nanometers (1 × 10 9). Thus, the result expressed in nanometers is divided by 1 × 10 9for conversion to meters.
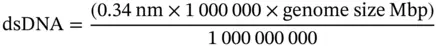
Depending on the organism, cells of different tissues can be characterized based on the number of sets of chromosomes present: monoploid (one set of chromosomes), diploid (two sets), triploid (three sets), tetraploid (four sets), pentaploid (five sets), and so on. For instance, the human genome contains 3.1 Gbp (3100 Mbp). Thus, in a human haploid (or monoploid) cell (e.g. a single set of chromosomes found in a gamete), the unfolded length of a single set of chromosomes, arranged linearly one after the other, would show an approximate length of:
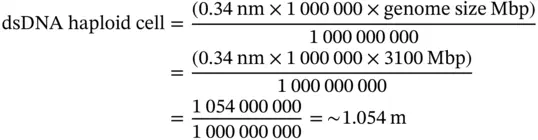

Thus, a single set of human chromosomes ( n = 23 Chr) can theoretically unfold up to 1 m. However, the human body is constituted mainly of somatic cells (diploid cells – two sets of chromosomes/cell). For a diploid cell (2 n = 46 Chr), the linear length of all 46 dsDNA molecules is calculated as above and the result in multiplied by two:
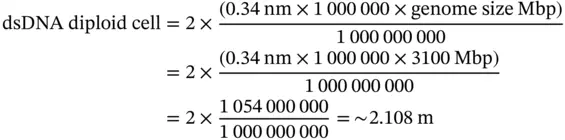

Therefore, the two sets (2 n = 46 Chr) of human chromosomes found inside a somatic cell can theoretically unfold up to 2.1 m. The linear length of dsDNA molecules from all chromosomes of a somatic cell and the estimated average number of somatic cells in the human body, can be used for various mental experiments (e.g. comparisons between DNA lengths and cosmic distances). These calculations can be empirically extended for ssDNA molecules placed linearly one after the other. For instance, the 2.1 m of dsDNA from a somatic cell, of course, doubles if the ssDNA approach is considered (2.1 m × 2 DNA strands = 4.2 m of ssDNA). The implementation found in Additional algorithm 2.1uses the above formula to convert the number of bases of a genome to physical length expressed in meters. Important: For convenience, from this point on all notations “b”, “kb”, “Mb”, “Gb” will refer to dsDNA (double stranded DNA).
Additional algorithm 2.1Note that the source code is in context and works with copy/paste.
document.write('Homo sapiens (3100 Mb):
'); document.write('DNA in a haploid cell nucleus: '); document.write(f(3100) + ' meters
'); document.write('DNA in a somatic cell nucleus: '); document.write((2 * f(3100)) + ' meters
'); function f(Mb){return (0.34 * 1000000 * Mb)/1000000000;} Output: Homo sapiens (3100 Mb): DNA in a haploid cell nucleus: 1.054 meters DNA in a somatic cell nucleus: 2.108 meters
Above, the example is given on Homo sapiens and the result shows the calculated total length of unfolded chromosomes for both haploid cells and diploid (somatic) cells. This computation can be applied to all genomes mentioned so far by calling function f repeatedly. Thus, Additional algorithm 2.1is extended to perform this calculation for an arbitrary number of species ( Additional algorithm 2.2).
Additional algorithm 2.2Note that the source code is in context and works with copy/paste.
// DNA to meters var a = 'Ambystoma mexicanum|32396Mb' + 'Pinus lambertiana|27603Mb' + 'Sequoia sempervirens|26537Mb' + 'Minicystis rosea|16Mb' + 'Sorangium cellulosum So0157-2|14.78Mb' + 'Escherichia coli|4.9Mb' + 'Encephalitozoon intestinalis|2.3Mb' + 'Ostreococcus tauri|12.6Mb' + 'Homo sapiens|3100Mb'; var t = a.split('Mb'); for (var u=0; u