Genomic and Epigenomic Biomarkers of Toxicology and Disease
Здесь есть возможность читать онлайн «Genomic and Epigenomic Biomarkers of Toxicology and Disease» — ознакомительный отрывок электронной книги совершенно бесплатно, а после прочтения отрывка купить полную версию. В некоторых случаях можно слушать аудио, скачать через торрент в формате fb2 и присутствует краткое содержание. Жанр: unrecognised, на английском языке. Описание произведения, (предисловие) а так же отзывы посетителей доступны на портале библиотеки ЛибКат.
- Название:Genomic and Epigenomic Biomarkers of Toxicology and Disease
- Автор:
- Жанр:
- Год:неизвестен
- ISBN:нет данных
- Рейтинг книги:4 / 5. Голосов: 1
-
Избранное:Добавить в избранное
- Отзывы:
-
Ваша оценка:
- 80
- 1
- 2
- 3
- 4
- 5
Genomic and Epigenomic Biomarkers of Toxicology and Disease: краткое содержание, описание и аннотация
Предлагаем к чтению аннотацию, описание, краткое содержание или предисловие (зависит от того, что написал сам автор книги «Genomic and Epigenomic Biomarkers of Toxicology and Disease»). Если вы не нашли необходимую информацию о книге — напишите в комментариях, мы постараемся отыскать её.
The latest developments in biomarker research applicable to toxicology and medicine Genomic and Epigenomic Biomarkers of Toxicology and Disease: Clinical and Therapeutic Actions
Genomic and Epigenomic Biomarkers of Toxicology and Disease: Clinical and Therapeutic Actions
Genomic and Epigenomic Biomarkers of Toxicology and Disease — читать онлайн ознакомительный отрывок
Ниже представлен текст книги, разбитый по страницам. Система сохранения места последней прочитанной страницы, позволяет с удобством читать онлайн бесплатно книгу «Genomic and Epigenomic Biomarkers of Toxicology and Disease», без необходимости каждый раз заново искать на чём Вы остановились. Поставьте закладку, и сможете в любой момент перейти на страницу, на которой закончили чтение.
Интервал:
Закладка:
Packaging into Vesicles and Mechanisms of Release
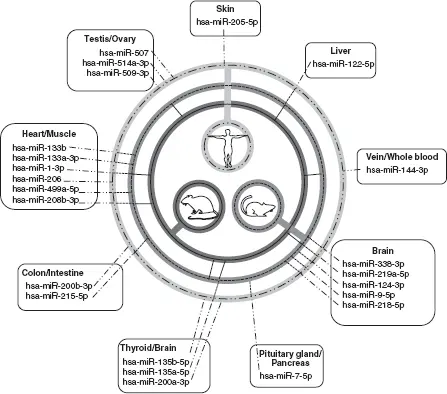
Figure 2.1 Tissue-specific miRNAs in mammals.Human tissue-specific miRNAs that were described as part of the human miRNA tissue atlas study (Ludwig et al. 2016) (light grey circle with dotted/dashed line) were cross-referenced with miRNA atlas studies that examine tissue-specific miRNAs in mouse (medium grey circle with dashed line) and rat (dark grey circle with solid line). Those miRNAs that transverse all three circles are ideal cross-species biomarkers for toxicological studies.
Both prokaryotic and eukaryotic cells release membrane-enclosed microvesicles that package a variety of different cellular-derived components, including nucleic acids, proteins, and lipids. The vesicles range in size (from small 50–150 nm exosomes to large 1,000–10,000 nm tumor cell-derived oncosomes), location of biogenesis, and content. These vesicles play important roles in cellular communications and their content is enriched in small non-coding RNAs (such as miRNAs, lncRNAs, snoRNAs, piRNA, snRNAs, etc.), but also in intact mRNAs and in fragmented tRNAs, mRNAs, lncRNAs, rRNAs, and other nucleic acids (Turchinovich et al. 2019). The roles of these packaged contents are not entirely clear for all RNA species; however, many studies have demonstrated that some of these RNAs can mediate differential responses in recipient cells. Many studies have focused on miRNAs in this paracrine role (reviewed in O’Brien et al. 2020) which therefore may make them serve as putative biomarkers of “active” release due to homeostatic, responsive, and perturbed cellular states. A number of characterization studies have been performed to determine the RNA contents of extracellular microvesicles isolated from human plasma, saliva, and urine, as well as cell lines of different lineages (see reviews in Amorim et al. 2017 and Turchinovich et al. 2019). In one of the first studies that use deep sequencing to profile extracellular vesicle (EV) RNAs, small RNAs were reported to dominate these “shuttle”-derived fractions; however, the distribution was different from that of the parent mouse dendritic cells co-cultured with cognate T cells (Nolte-’t Hoen et al. 2012). Importantly, whereas miRNAs dominated the intracellular small RNA content (~55% of the reads), miRNAs were only the fourth most populated of the EV-derived small RNAs, at around 10% of the total reads. Later studies have both supported and contradicted these findings. For example, Sork et al. (2018) also described different patterns of intracellular small, ribosomal, and other RNAs versus those found in EVs. Of the total reads, small RNAs were a minority; they were noted in the five cell cultures assessed, ranging from ~ 2% to ~ 40% of the reads. However, 58%–83% of the small RNA EV reads were attributed to miRNAs, many of these reflecting those found in the parent cell, with some notable exceptions such as miR-451a. The disagreements between different studies could be due to cell types, methods, and the analyses used. Overall, from a biomarker perspective, the content of these EVs reflects the cellular state and can be specific to a cell type by partially reflecting the transcriptome (Sork et al. 2018; Srinivasan et al. 2019); yet the EVs differ in intracellular RNA content from the source cell (Guduric-Fuchs et al. 2012; Zhang et al. 2010). Therefore packaging these EVs is not merely representative of the RNA cellular milieu; it is rather an active process that may be influenced by cellular responses to stressors.
Many recent studies have focused on determining mechanisms that are involved in EV formation and RNA packaging. The initial formation varies for the different types of EVs. For example, intraluminal vesicles form through the invagination of endosomes known as multivesicular bodies (MVBs), which are then guided by cytoskeletal components and degraded by giant intracellular lysosomes or released into the extracellular space after fusion with the plasma membrane (see reviews in Janas et al. 2015 and O’Brien et al. 2020). Other EVs, such as microvesicles, larger oncosomes, and apoptotic bodies, form directly from the plasma membrane itself. During these processes, RNAs are selectively packaged by the vesiculation machinery through different interactions. These vesicles are enriched with cholesterol, sphingomyelin, glycosphingolipids, and phosphatidylcholine comprised of saturated fatty acids when they are derived from raft-like regions of the MVBs and the plasma membrane. Ceramides, which preferentially locate on the outside rafts of MVBs, are thought to associate with ribonucleoprotein (RNP) complexes and may help selectively package RNA into EVs (Kosaka et al. 2013). Both RNA sequence and RNA secondary structure can mediate this selection (Janas et al. 2006). The sumoylated heterogenous nuclear RNP (specifically, hnRNPA2B1) can recognize the GGAG motif in miRNAs (e.g., miR-198 and -601) and subsequently transfer into exosomes (Villarroya-Beltri et al. 2013) without itself being packaged (Zhou et al. 2020). Sumoylated hnRNPA1 may work in a similar fashion (Li et al. 2004). Also, the synaptotagmin-binding cytoplasmic RNA-interacting protein (SYNCRIP) mediated the sorting of miRNAs bearing the GGCU motif in hepatocyte cell culture (Santangelo et al. 2016). These seed sequences, also termed “EXO-motifs,” are located in the 3ʹ sites of miRNAs (Janas et al. 2006), whereas the sites that mediate mRNA targeting (and primary function) are located in the 5ʹ end. This suggests that the loading of miRNAs into vesicles could occur even if there were very different mRNA targeting functions. It has also been noted in B cells that the post-transcriptional addition of uracil at the 3ʹ end of the miRNA sequence preferentially sorts into exosomes, whereas adenylated miRNAs remain cell-bound (Koppers-Lalic et al. 2014). Other RNA-binding proteins implicated in preferential RNA loading include AGO2 (McKenzie et al. 2016), annexin A2 (Hagiwara et al. 2015), major vault protein (Statello et al. 2018), YBX1 (Kossinova et al. 2017; Shurtleff et al. 2016; Yanshina et al. 2018), lupus La protein (Temoche-Diaz et al. 2019), and Arc1 (Ashley et al. 2018) (see review in O’Brien et al. 2020).
Biological Interpretations Due to Mechanisms of Release
The difference between a “leakage” miRNA biomarker and one that is non-randomly packaged into an EV as a putative communication signal, is an important distinction for development of miRNA biomarkers that are indicative of health effects. For example, mammalian miR-122 has been heavily investigated as a leakage biomarker for liver toxicity, in the context of both non-clinical and clinical species. miR-122 accounts for approximately 72% of the total liver miRNAs in mice (Lagos-Quintana et al. 2002) and is highly specific for human, mouse, and rat livers (Landgraf et al. 2007; Smith et al. 2016). In a landmark study by Wang et al. (2009), mice were treated with hepatotoxic levels of acetominophen (APAP) and miRNA microarrays were used to measure difference in the plasma. Forty-four miRNAs were significantly altered, miR-122 showing a 470-fold increase twenty-four hours after APAP overdose, and these alterations could be observed as soon as one hour after exposure. This APAP overdose or acute liver injury-associated miR-122 release was later confirmed in human patients (Starkey Lewis et al. 2011). The observation that decreased liver-based miRNAs with APAP hepatotoxicity correlated with increases in serum plasma suggested that these miRNAs were released as packaged contents or leaked during hepatocyte necrosis; the leakage theory was supported by Arroyo et al. (2011), who demonstrated that miR-122 was primarily associated with a RNP complex (Ago2) and was not membrane-bound as in exosomes. However, later studies provided clues that the mechanism of miR-122 release from liver was context-dependent, that is, dependent on factors such as the condition of the liver, the timing of the injury or perturbation, the half-life of a miRNA, and baseline variability. Subtoxic exposures in vitro result primarily in hepatocyte-derived release of exosome-bound miR-122 (Holman et al. 2016; Mosedale et al. 2018), and alcoholic and inflammatory liver disease was associated with exosome-bound miR-122—as opposed to drug-induced injury, which was found in the protein-bound fraction of plasma (Bala et al. 2012). Further, the protection of miRNA from RNases present in biofluids may preferentially enrich exosome and protein-bound miRNAs (Arroyo et al. 2011; Koberle et al. 2013; Li et al. 2012; Turchinovich et al. 2011).
Читать дальшеИнтервал:
Закладка:
Похожие книги на «Genomic and Epigenomic Biomarkers of Toxicology and Disease»
Представляем Вашему вниманию похожие книги на «Genomic and Epigenomic Biomarkers of Toxicology and Disease» списком для выбора. Мы отобрали схожую по названию и смыслу литературу в надежде предоставить читателям больше вариантов отыскать новые, интересные, ещё непрочитанные произведения.
Обсуждение, отзывы о книге «Genomic and Epigenomic Biomarkers of Toxicology and Disease» и просто собственные мнения читателей. Оставьте ваши комментарии, напишите, что Вы думаете о произведении, его смысле или главных героях. Укажите что конкретно понравилось, а что нет, и почему Вы так считаете.












![John Bruce - The Lettsomian Lectures on Diseases and Disorders of the Heart and Arteries in Middle and Advanced Life [1900-1901]](/books/749387/john-bruce-the-lettsomian-lectures-on-diseases-and-disorders-of-the-heart-and-arteries-in-middle-and-advanced-life-1900-1901-thumb.webp)