Each and every organism has a different genetic (DNA) makeup, which in turn makes the whole organism different with respect to their carbohydrates, lipids, and proteins. This is due to the fact that DNA transcribes and translates to mRNA and proteins, respectively (central dogma). This makes DNA the choice for manipulation in genetic engineering as manipulating it leads to the generation of a whole new organism. This postulation gives rise to many other disciplines of genetic engineering like recombinant protein production, protein engineering, metabolic engineering, etc.
Every organism being different makes it difficult to use proteins and other biomolecules of one organism to the other. This was the main reason why proteins/enzymes from animals cannot be used by humans. Earlier, insulin was extracted from the pancreas of slaughtered pigs, posing a threat to human health. This leads to the discovery of the first recombinant product, Insulin, approved by the US Food and Drug Administration (FDA) in 1982 (Goeddel et al., 1979). Now synthetic insulin is easily being produced by yeast worldwide as Escherichia coli does not perform post‐translational modifications required to form functional insulin.
Similarly, genetic engineering is now used to produce several other biocommodities. Modifying DNA and getting it expressed inside the host organism requires several steps, as shown in Figure 1.1and the number of enzymes. These enzymes are explained in further sections with other requirements for genetic engineering.
1.2.1 DNA‐altering Enzymes
The basis of rDT is the manipulation of DNA molecules with the help of molecular biology tools and biocatalysts. The available purified enzymes that can manipulate DNA molecules with specific changes can be categorized in four broad classes: (i) DNA polymerases, (ii) nucleases, (iii) DNA ligases, and (iv) end‐modification enzymes.
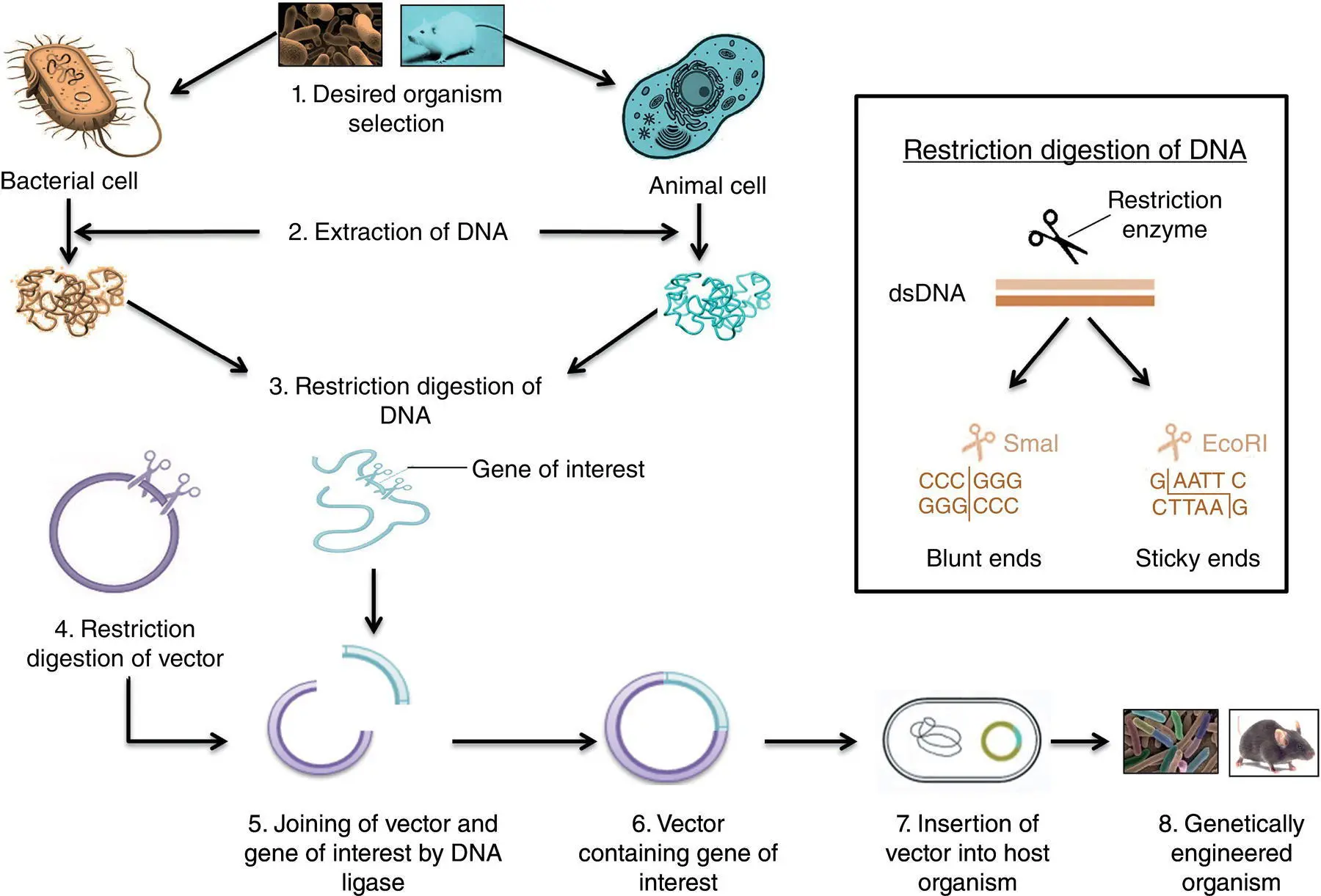
Figure 1.1 Basic steps of gene cloning.
DNA polymerases is the key enzyme in DNA replication driving the synthesis of new DNA strand from the parent DNA or RNA strand acting as a template. DNA polymerases require an oligo nucleotide (primer) for the initiation of DNA strand synthesis. DNA polymerase‐I (DNA‐dependent DNA polymerase) is widely studied polymerase and has both polymerization and exonuclease activity that can help in synthesizing new strand as well as the degradation for proof reading or repair and primer removal. DNA polymerase I (or Pol I) takes part in the process of prokaryotic DNA replication. It was the first DNA polymerase discovered by Arthur Kornberg in 1956 (Lehman et al., 1958). Pol I has three different enzymatic activities: A 5′ →3′ DNA‐dependent DNA polymerase activity, a 3′ →5′ exonuclease activity that helps in proofreading, and a 5′ → 3′ exonuclease activity mediating nick translation during DNA repair. Pol I having polymerase but lacking nuclease activity is called klenow fragments (Klenow and Henningsen, 1970; Jacobsen et al., 1974).
Taq DNA polymerase, a thermostable DNA polymerase isolated from Thermus aquaticus by Chien et al. (1976). It is frequently used in the polymerase chain reaction (PCR), to amplify small quantities of DNA. It has a functional 5′ → 3′ exonuclease domain at the N‐terminal, and 3′–5′ exonuclease domain was changed so it is not functional. Optimum temperature for Taq pol activity is 75–80 °C, with a half‐life of greater than 2 hours at 92.5 °C and minimum 9 minutes at 97.5 °C, and able to replicate a 1000 bp strand of DNA within 10 seconds at 72 °C.
Nucleases can cut or digest the DNA molecules either from one end or in middle by acting on phosphodiester bond, which forms the backbone of DNA (Nishino and Morikawa, 2002). Depending on the position of their digestion nucleases are of two types: exonucleases and endonucleases. Phosphodiester bonds present in the ends of the DNA are digested by exonucleases, removing nucleotides one at a time from either end. Whereas phosphodiester bonds present in the middle of the DNA strand are digested by endonucleases. Specificity of nucleases vary from source to source, Aspergillus oryzae ’ s endonuclease only cleaves single strands, whereas deoxyribonuclease I (DNase I), extracted from cow pancreas, cuts single as well as double‐stranded DNA molecules. DNase I not being sequence specific cuts DNA at random interior phosphodiester bond, leading to production of mononucleotides and very short oligonucleotides mixture. Some of the examples of nucleases are (i) Mung Bean Nuclease (isolated from mung bean sprouts) – a single‐strand‐specific DNA and RNA endonuclease which can degrade single strand overhangs from the end of DNA and RNA to make blunt ends. (ii) Nuclease S1 (isolated from Aspergillus sp.) – S1 nuclease is a single‐strand‐specific endonuclease that hydrolyzes single‐stranded RNA or DNA into 5′ mononucleotides. The enzyme will hydrolyze single‐stranded extensions in duplex DNA such as loops and gaps. S1 Nuclease is stable at 65 °C (Balabanova et al., 2012). (iii) Exonuclease III (isolated from E. coli ) – removes single nucleotides from 3′ termini of the duplex DNA. It is generally used to make a set of nested deletions of the terminal of linear DNA strand. (iv) BAL31 nuclease (isolated from Alteromonas espejiana ) – it is a 3′‐exonuclease and removes nucleotides from both 3′‐terminus of the two strands of linear DNA. (v) RNase H – it is an endonuclease that specifically hydrolyzes the phosphodiester bonds of RNA, when the RNA is hybridized to DNA (RNA‐DNA) (Cerritelli and Crouch, 2009). (vi) RNase P – the specificity of this enzyme is to cleave other RNA molecules at the junction of single‐stranded and the 5′ end of double‐stranded regions of RNA (Guerrier‐Takada et al., 1983).
Restriction endonuclease (RE) enzymes recognize and cleave the specific phosphodiester bond present in the DNA molecule (Smith and Welcox, 1970). Restriction enzymes are broadly classified into Type‐I, Type‐II, and Type‐III. For their functioning, they require specific temperature, ATP, and divalent magnesium ions. On digestion of the DNA molecule they can produce both blunt and sticky end. Type I REs interact with unmodified target site in dsDNA. They are bifunctional enzymes having methylase and endonuclease in a single protein molecule. They cleave DNA around 1000 bp away from the recognition site. For their function, both ATP and Mg 2+are required. Type II REs are highly specific and cleave within or very near to the recognition sequence due to this reason type II are used widely in genetic engineering. They do not require ATP for the restriction digestion, only Mg 2+is required. Type III REs cleave dsDNA at defined positions and need ATP, Mg 2+. They cleave the DNA 24–26 bp away from the specific site.
Ligases are considered to be molecular glue in rDT, used to join two DNA segments. It also has role in various aspects of molecular biology such as replication, recombination, and cloning. In the presence of ligase enzyme when two DNA fragments are mixed under a certain condition, base pairing between two fragments occur which results in sealing of two different DNA fragments to make a chimera (Pascal et al., 2004). It occurs due to covalent bonds formation between 2′‐PO 4group and 3′‐OH group of adjustment strands.
1.2.1.3.1 Mechanism of Action
The DNA ligation reaction has two main steps. First, the DNA ends collide with each other by chance and reside together for enough time for the ligase to join them. For sticky overhangs, there is an additional reason; lower temperature stabilizes the hydrogen bonding between the complementary nucleotides, which really helps to join easily. DNA ligase catalyzes the joining of the 3′‐OH to the 5′‐phosphate via a two‐step process. First, the AMP nucleotide, which is linked to a lysine residue in the enzyme’s active site, is shifted to the 5′‐phosphate. Then the AMP‐phosphate bond is attacked by the 3′‐OH, forming the covalent bond and discharging AMP. To allow the enzyme to conduct further reactions, the AMP in the enzyme’s active site must be restocked by ATP. The DNA ligase enzyme has optimal activity at 25 °C so the ligation reaction is carried out at a temperature range of 16–25 °C. Normally, 1 hour at 16 °C is best for ligation but since bringing the DNA ends together is the least efficient part of the reaction favoring this by lowering the temperature to 4 °C can give even more efficiency. Two types of ligases are there that are used in genetic engineering. E. coli ligases which is generally used to join two sticky or cohesive ends. It is mainly used to catalyze a phosphodiester bond between duplex DNA‐containing cohesive ends. It will not efficiently ligate blunt‐ended fragments as much as it efficient for sticky end ligation. NAD +is required as a cofactor for the ligation reaction. T4 DNA ligase that is generally used for the ligation of blunt‐ended DNA molecules. This enzyme can join blunt‐ended termini as well as ends with cohesive overhangs (either 3′ or 5′ complementary overhangs). This enzyme will also function for the repair of single‐stranded nicks in duplex DNA, RNA, or DNA/RNA duplexes. ATP is required as a cofactor for the ligation reaction. All the reactions are generally carried out at 16 °C. Sometimes PEG (polyethylene glycol) can be added that helps in fusion and increases the frequency of collision between two DNA molecules.
Читать дальше













